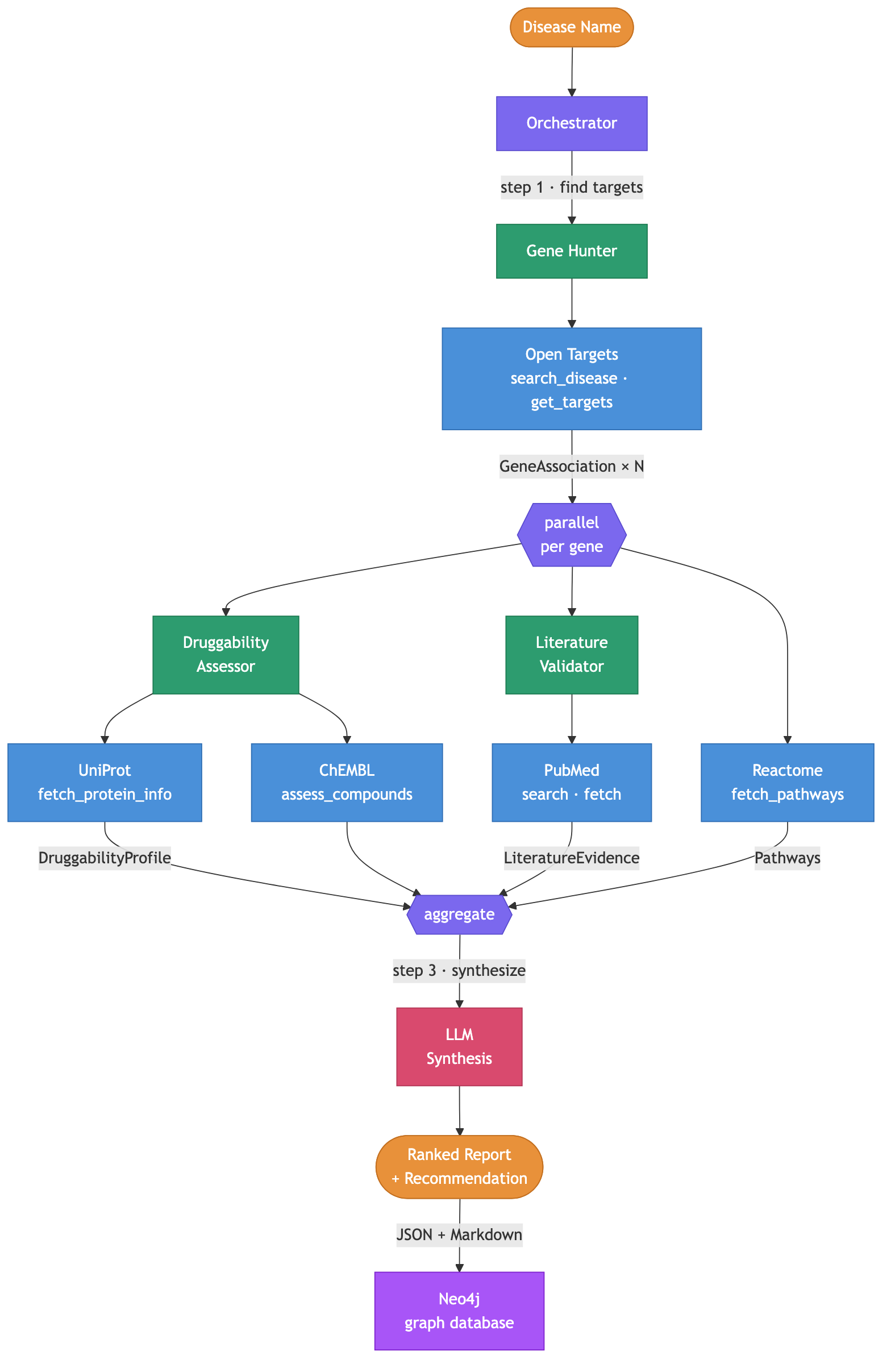
Building a multi-agent system for drug target discovery
Update (2026-03-24): Since the original post, I added three things. First, a Reactome pathway tool that fetches biological pathways for each gene target. Second, a Neo4j graph database for accumulating results across pipeline runs. Reports are ingested as a knowledge graph (diseases, genes, proteins, compounds, papers, pathways) that enables cross-disease queries like “which targets overlap between Alzheimer’s and Parkinson’s?” Third, a web frontend at bio.arcosdiaz.com for interactive exploration of the graph (still work in progress). All three are described below. ...
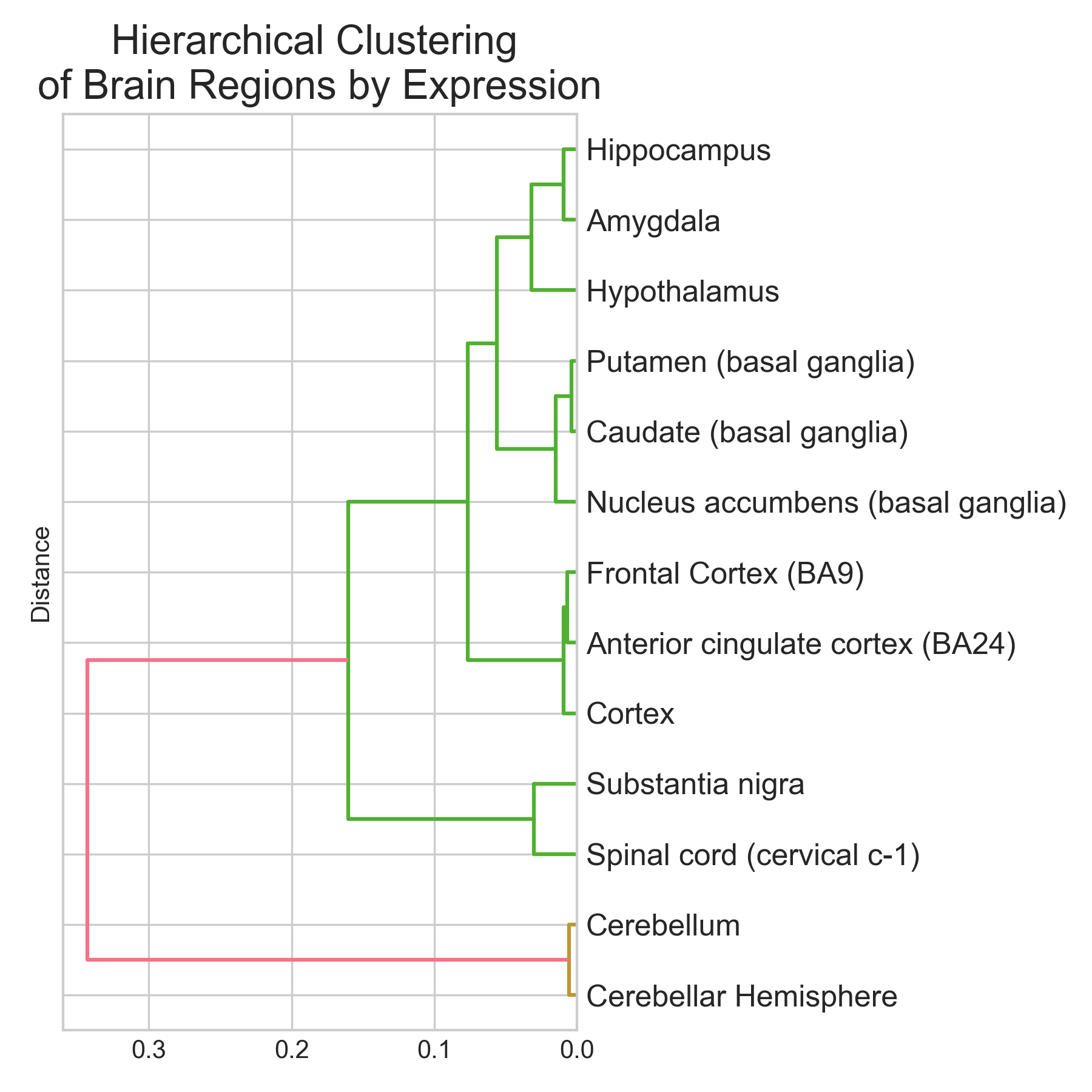
Classifying brain regions from gene expression RNA-seq data
tl;dr I trained three classifiers (Logistic Regression, Random Forest, XGBoost) to predict brain region of origin from GTEx bulk RNA-seq expression profiles across 13 brain regions and 2,642 samples. XGBoost did best: 95.1% accuracy (5-fold CV: 94.9 +/- 0.9%), macro-averaged AUROC near 0.99. Cerebellum and spinal cord were classified perfectly (F1 = 1.00). Basal ganglia subregions (caudate, putamen, nucleus accumbens) were hardest to separate (F1 ~ 0.89-0.96), which makes sense given their shared developmental origin. The top discriminative genes are not statistical artefacts. They map onto known neurobiology: RORB (#2, cortical layer IV marker), GAL and TRH (#9 and #19, hypothalamic neuropeptides), and a cluster of cerebellar-specific genes (ARHGEF33, HR, KCNJ6) all appear near the top. Non-coding RNAs (lncRNAs + pseudogenes) make up ~37% of the top 30 features. The brain has the highest proportion of non-coding transcription of any organ, so this isn’t surprising. Disclaimer: This was a hobby project. I tried to be rigorous, but these results are an initial exploration, not an exhaustive analysis. The pseudogene hits at the top of the ranking especially need validation to rule out mapping artefacts. ...

Graph Convolutional Networks for Fraud Detection of Bitcoin Transactions
tl;dr I trained 4 different types of models to classify bitcoin transactions. For each, two versions of the feature set were used: all features (local + neighborhood-aggregated) and only local features (without neighborhood information). The best model was a Random Forest trained with all features: its performance was impaired when the aggregated features were removed. The best graph-based neural network model was APPNP and its performance was better when only local features were used. APPNP performed better than an MLP with comparable complexity, indicating that the graph structure information gave it an advantage. Finally, the best GCN model required using all features and several strategies to reduce overfitting. The excellent performance of a Random Forest shows that it makes sense to consider simple models when faced with a new task. It also indicates that the individual node features in the Elliptic dataset are already informative enough to make good predictions. It would be interesting to explore how the model performs, when fewer samples and/or features are available for training. ...

Personalized Medicine Kaggle Competition
This notebook describes my approach to the Kaggle competition named in the title. This was a research competition at Kaggle in cooperation with the Memorial Sloan Kettering Cancer Center (MSKCC). The goal of the competition was to create a machine learning algorithm that can classify genetic variations that are present in cancer cells. Tumors contain cells with many different abnormal mutations in their DNA: some of these mutations are the drivers of tumor growth, whereas others are neutral and considered passengers. Normally, mutations are manually classified into different categories after literature review by clinicians. The dataset made available for this competition contains mutations that have been manually anotated into 9 different categories. The goal is to predict the correct category of mutations in the test set. ...