I built a multi-agent system in plain python that takes a disease name and autonomously finds potential drug targets by querying public bioinformatics databases. You enter “Alzheimer disease” as an input and returns a ranked list of targets, each annotated with protein structure data, known compounds, clinical trial progress, and recent literature. I ran it on three diseases and the results matched real-world pharma consensus in every case, without any hardcoded domain knowledge.
For Alzheimer’s, the system identified APP as the top target (55 known compounds, Phase 3 trials) and flagged APOE as the strongest genetic risk factor but undruggable: zero compounds. For Parkinson’s, LRRK2 came out first, which is the kinase that Denali and Biogen are both currently targeting using clinical-stage inhibitors. For schizophrenia, DRD2, because every antipsychotic on the market targets the dopamine D2 receptor.
The problem
Early-stage drug target identification is basically a cross-referencing exercise. Open Targets tells you which genes are associated with a disease. UniProt tells you what the protein looks like: its family, subcellular location, whether it has a solved 3D structure. ChEMBL tells you if compounds exist that bind it, and how far they have progressed in trials. PubMed tells you what the literature says about the gene-disease association.
A scientist doing this manually queries each database, copies results into a spreadsheet, and writes a recommendation. I wanted to know if an LLM-coordinated agent system could do this end-to-end and produce results that are biologically valid.
Architecture
The system uses three specialized agents and one orchestrator.
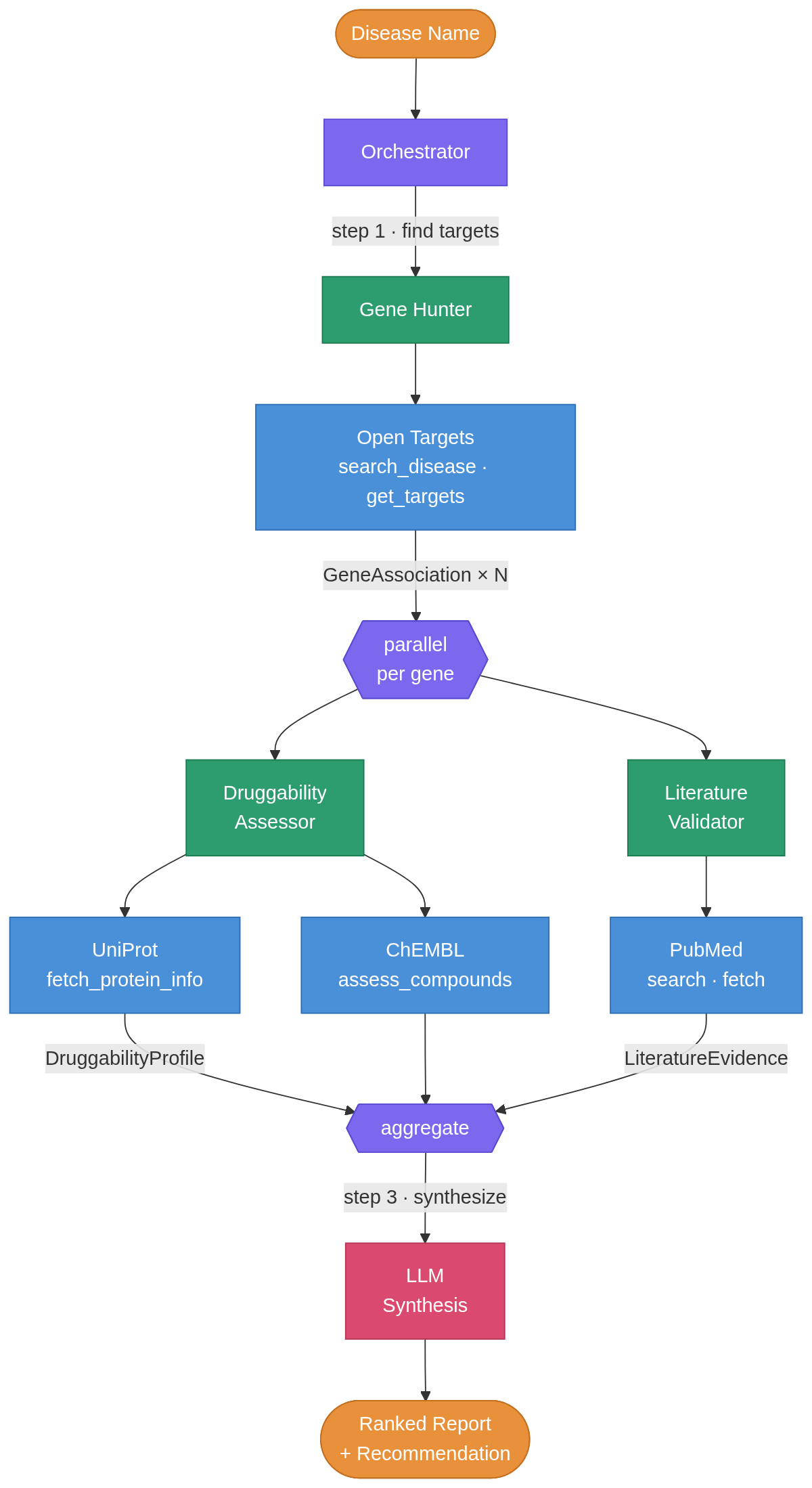
The Gene Hunter queries Open Targets’ GraphQL API for disease-gene associations and returns the top-ranked genes. Then two agents run in parallel via asyncio.gather() for each gene: the Druggability Assessor hits UniProt for protein annotations and ChEMBL for compound/bioactivity data, while the Literature Validator searches PubMed and gets recent abstracts. Each agent uses Gemini 2.5-flash (free tier, but you can use more advanced models) for its reasoning step, interpreting raw protein data into a druggability verdict or classifying literature evidence as supporting, contradicting, or inconclusive.
The orchestrator then compiles everything into a ranked report plus recommendation.
I separated the agents because the required reasoning is different. The Druggability Assessor interprets protein families and compound binding data (pharmacology). The Literature Validator reads abstracts and weighs conflicting evidence (biomedical text analysis). Putting both in one prompt would make it less specific. The architecture is also modular: adding a Clinical Trials agent, Pathway Analysis agent, and others, would not require touching existing code.
I deliberately avoided LangGraph and similar agent frameworks. The orchestration logic is just async Python: a few gather() calls and some loops. Pydantic models define the data contracts between agents. I might extend it to add functionality that really requires a such a framework.
What it found
Alzheimer disease
The system ranked five targets. APP (amyloid beta precursor protein) came out on top: 55 known compounds, Phase 3 trials, direct causal role in early-onset AD through the amyloid pathway. APP has been the central therapeutic hypothesis in Alzheimer’s for decades, so this is what you would want to see.
APOE is the more interesting result. The system flagged it as the strongest genetic risk factor for late-onset AD but noted zero compounds and zero clinical progress. APOE is a lipid transport protein, and the field has been trying to figure out how to drug it with small molecules for years without success. The system identified that gap from the data alone.
PSEN1 and PSEN2 were identified as gamma-secretase components (peptidase A22A family) with no clinical-phase compounds, and the system flagged toxicity concerns. This maps onto real history: gamma-secretase inhibitors like semagacestat and avagacestat failed in trials because of toxicity from Notch signaling disruption. The database queries can’t directly surface those specific failures, but the protein annotations were enough for the LLM to flag the risk.
Parkinson disease
LRRK2 ranked first: a protein kinase with 54 known compounds and Phase 4 annotation. It is the most actively pursued kinase target in Parkinson’s right now, with inhibitors from Denali and Biogen in clinical trials.
SNCA (alpha-synuclein) was noted as “intrinsically disordered,” meaning it does not have a stable, well-defined 3D structure under physiological conditions and is challenging for conventional small molecule drug design, even though it is central to PD pathology. The system understood that distinction, which is the kind of thing I was hoping it could do: not just retrieve data, but reason about what makes a protein a good target.
PRKN (Parkin), an E3 ubiquitin ligase with zero compounds, was flagged as a candidate for PROTAC-type approaches (therapeutic effect by inducing degradation of the protein rather than inhibiting its function). Its partner kinase PINK1 was identified with activators entering trials. Both are in the mitophagy pathway (removal of damaged mitochondria), and both calls are reasonable.
Schizophrenia
DRD2: Phase 4 (approved drugs), G-protein coupled receptor, 57 compounds, strong genome-wide association study evidence. Every antipsychotic targets it. Not surprising, so the system should find it.
SHANK3 was flagged as an intracellular scaffold protein with zero compounds. It is hard to target a protein-protein interaction hub sitting inside the postsynaptic density, and the system said as much.
DRD3 showed 99 compounds at Phase 2. For example, cariprazine: a DRD3-preferring partial agonist approved for schizophrenia.

Under the hood
The pipeline is fully async. Each gene’s druggability and literature assessments run concurrently, with staggered delays for PubMed’s rate limits (3 retries, 2-second backoff on 429 responses).
Each agent has a separate call_llm() function, which makes testing simple: the test files mock all HTTP calls with respx and all LLM calls with AsyncMock. No live API calls required in the test suite.
Five Pydantic models define the contracts between agents: GeneAssociation, DruggabilityProfile, LiteratureEvidence, TargetReport, ReconReport. The orchestrator composes them into a final report serialized as both JSON (for downstream analysis) and Markdown (for reading).
All four APIs are free. Open Targets, UniProt, and ChEMBL need no authentication. PubMed just wants an email address.
Limitations
The system queries only Open Targets for gene-disease associations. GWAS Catalog or DisGeNET would improve coverage. PubMed returns abstracts, not full text, so the literature agent misses nuance in methods sections and supplementary data. The clinical trial integration is limited to ChEMBL’s “max phase” field information: a ClinicalTrials.gov query agent would add real trial design details.
The LLM evidence classifications are sometimes too cautious. Several targets got “inconclusive” when the literature clearly supports their role. But this is a prompt engineering problem, not an architectural one.
There’s also no cross-disease analysis. The system cannot answer “which targets overlap between Alzheimer’s and Parkinson’s?” since each run is independent.
What I’d build next
A graph database (Neo4j or Kuzu) for accumulating results across queries. Right now each run is isolated, and there are interesting questions about what is shared between diseases. I would also want a Clinical Trials agent for ClinicalTrials.gov data and a Pathway Analysis agent using Reactome or KEGG, because treating each gene in isolation misses half the biology.