Classifying brain regions from gene expression RNA-seq data
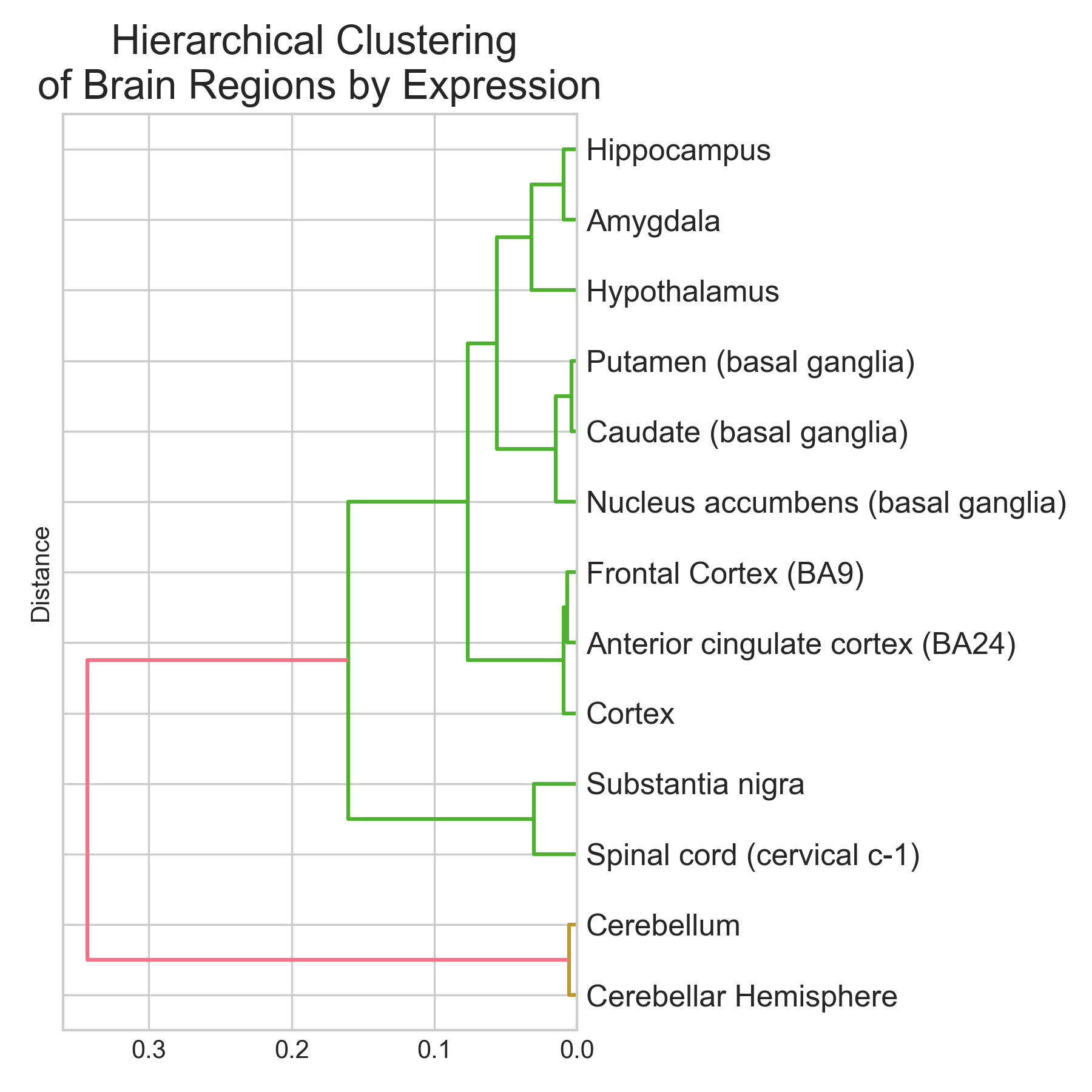
tl;dr I trained three classifiers (Logistic Regression, Random Forest, XGBoost) to predict brain region of origin from GTEx bulk RNA-seq expression profiles across 13 brain regions and 2,642 samples. XGBoost did best: 95.1% accuracy (5-fold CV: 94.9 +/- 0.9%), macro-averaged AUROC near 0.99. Cerebellum and spinal cord were classified perfectly (F1 = 1.00). Basal ganglia subregions (caudate, putamen, nucleus accumbens) were hardest to separate (F1 ~ 0.89-0.96), which makes sense given their shared developmental origin. The top discriminative genes are not statistical artefacts. They map onto known neurobiology: RORB (#2, cortical layer IV marker), GAL and TRH (#9 and #19, hypothalamic neuropeptides), and a cluster of cerebellar-specific genes (ARHGEF33, HR, KCNJ6) all appear near the top. Non-coding RNAs (lncRNAs + pseudogenes) make up ~37% of the top 30 features. The brain has the highest proportion of non-coding transcription of any organ, so this isn’t surprising. Disclaimer: This was a hobby project. I tried to be rigorous, but these results are an initial exploration, not an exhaustive analysis. The pseudogene hits at the top of the ranking especially need validation to rule out mapping artefacts. ...
